Sno-Code
by Leif Hanlen
A Snomed CT Text Analysis suite
We’ve released our work on Snomed-CT text analysis, our first step is a public web service that can analyse English text and report any SNOMED CT concepts that are detected. The service is available at http://snomedct.t3as.org/
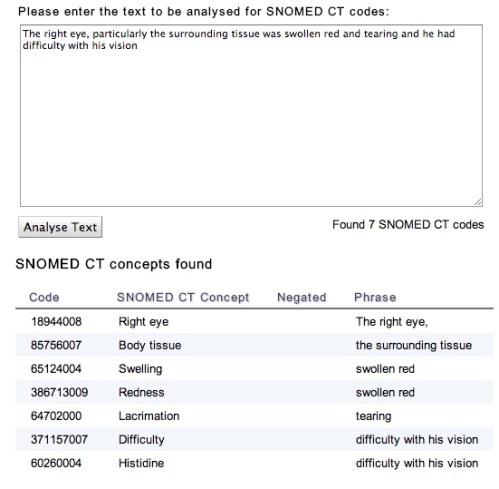
The service uses the National Library of Medicine’s software product MetaMap and the National Library of Medicine’s Unified Medical Language System (UMLS). The screenshot below is generated from the web page, using the input text of Figure 3, of Sager et al. Automatic encoding of Snomed III
If you would like to try out some other clinical texts, you might try the Medical History texts from Monash University.
Please note: This demonstration service runs on a publicly accessible server that is not geographically constrained. All text entered in the web page is sent in clear text to the Amazon service. Please do not upload private clinical documents.
The page is a simple front-end for the public web service that can analyse English text and report SNOMED CT concepts that are detected. At present we are using a heuristic to filter the semantic types that MetaMap returns to restrict the hits to clinical terms. We will improve the heuristic over time.
The next feature we add will be to also display the semantic type of each returned concept to feed into a system that allows the user to view/search-over specific types with some defaults set.
For developers, have a look at the service page to find out how to use the service as an API. We will shortly be releasing the source code as well.
SnoMed
SNOMED CT or SNOMED Clinical Terms is a systematically organized computer processable collection of medical terms providing codes, terms, synonyms and definitions used in clinical documentation and reporting. Wikipedia
More information on the Australian terminology SnoMed CT-AU can be found at the NEHTA site.
MetaMap
MetaMap is the most common tool in clinical text analysis applications to codify and relate medical concepts. We found only 1 hit on PubMed for the similar systems of Mgrap, and 12 hits for KnowledgeMap. There were 74 hits for MetaMap. MetaMap has resulted in substantial performance gains. For example,
- in biomedical paper search, MetaMap improves hit correctness by 15% (Aronson AR and Rindflesch TC. Query expansion using the UMLS Metathesaurus. Proc AMIA Annu Fall Symp. 1997:485–489,
- in biomedical paper summarisation, MetaMap makes text 83% more compact (Hofmann O, Schomburg D. Concept-based annotation of enzyme classes. Bioinformatics. 2005;21(9):2059–2066.)
With thanks to Mats Henrikson, Hanna Suominen and Neil Bacon